CRISPR-Cas silencing of Apol11b in Murine Erythroleukemia Cells

Students will contribute to research using CRISPR technology to create complete gene knockout lines, helping to clarify the exact function of the gene Apol11b.
This project has reached full capacity for the current term. Please check back next semester for updates.
Project description
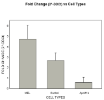
We have previously determined that the gene Apol11b is transiently expressed during the process of erythroid terminal differentiation. However, the exact function of this gene remains unknown. Recent experiments using interfering RNA suggest it may play a role in hemoglobin production, as in cells carrying a silencing construct there is a clear differences in hemoglobin accumulation compared to controls. The data concerning expression of Apol11b mRNA and protein have been inconsistent, possibly due to variability in the silencing of the gene by siRNA. We now will take the approach of creating complete gene knockout lines using CRISPR technology to generate more consistent and reproducible results.
Project outcome
Students will be expected to keep a detailed digital laboratory notebook on Dr. Koury's UB Box drive and present their results at regular lab meetings held with Dr. Koury and other graduate/undergraduate students in the lab. Data generated of sufficient quality will also be included in manuscripts/presentations at meetings with co-authorship for the student researchers. Data will also be presented at UB and possibly other regional student-centered scientific meetings in poster format in the spring semester.
Project details
| Length of commitment | About a semester (3-5 months) |
| Start time | Anytime |
| In-person, remote, or hybrid? | In-person |
| Level of collaboration | Small group project (2-3 students) |
| Benefits | Academic Credit |
| Who is eligible | Students with good basic laboratory skills, familiarity with basic laboratory instrumentation, pipetting, some cell culture and record keeping, hands-on experience with basic molecular biology techniques desired |
Project mentor
Stephen Koury
Research Associate Professor
Biotechnical and Clinical Laboratory Sciences
Start the project
- Email the project mentor using the contact information above to express your interest and get approval to work on the project. (Here are helpful tips on how to contact a project mentor.)
- After you receive approval from the mentor to start this project, click the button to start the digital badge. (Learn more about ELN's digital badge options.)
Preparation activities
Once you begin the digital badge series, you will have access to all the necessary activities and instructions. Your mentor has indicated they would like you to also complete the specific preparation activities below. Please reference this when you get to Step 2 of the Preparation Phase.
- Students who have taken MT437/537, Applications of Molecular Biotechnology, with Dr. Koury should review their notes and text entries related to CRISPR-Cas gene editing.
- All students should also read a free, full-text CRISPR-Cas9 review article available on PubMed: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6466564/.
Keywords
Biotechnical and Clinical Laboratory Sciences